Welcome to the ConesaLab website!
Latest papers and news
July: From Valencia’s pool to Liverpool
As summer has finally arrived, things in the ConesaLab have...

















Research Lines
Long-reads transcriptomics
We are developing methods and software for the analysis of alternative isoform expression and its effect on the phenotype.
Integration of Multi-Omics data
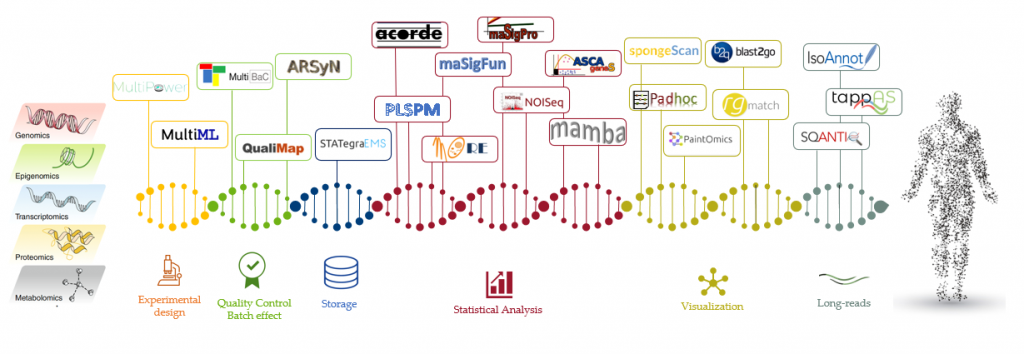
We have created a wide array of tools for the analysis of multi-omics data including experimental design, removal of batch effects, simulation of datasets, integration, and visualization.
Single-cell and spatial transcriptomics
We are developing cutting edge algorithms to analyze single cell data, taking special focus on the use of long read data. This goes in combination with spatial-transcriptomics, shaping the future of the field.
Resource and community building
We want to provide scientists easy and accessible tools for their research. Thus, we have created tutorials and guidelines to work with our tools.
Our Tools
Our lab develops statistical methods and user-friendly software tools to analyze these multi-omics data. Our most current interest is the application of long reads sequencing technologies for transcriptome analysis and the integration of multi-omics data to model chromatin-metabolome regulation.